Help
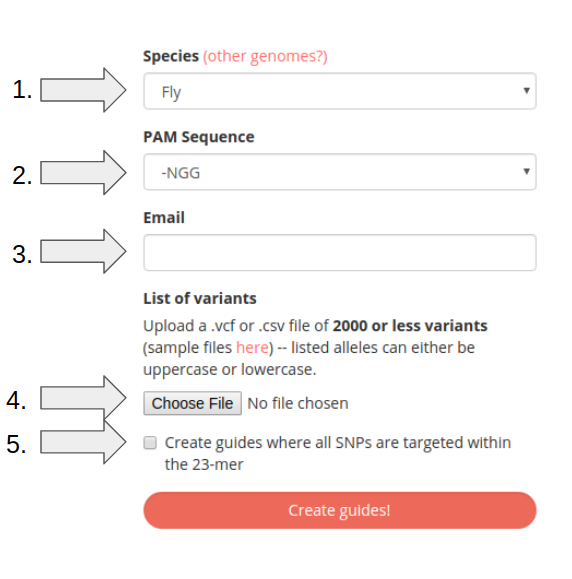
Step 1: Choose Species
Select the species of interest from the drop down menu. If the species you're interested in targeting is not listed please fill out the request form here.
Step 2: Choose PAM Sequence
Select the PAM sequence from the second drop down.
-NGG and -NAG sequences supported.
Step 3: Enter Email Address
Enter your email address. It is necessary to provide an email as a link to the design results will be sent to the provided email.
Step 4: Upload Variant Information
Click the choose file button, then select the .vcf or .csv file containing the variants to be targeted. Both Variant Call Format (VCF) and a simplified csv format described here are supported. For more information on VCF format please see the following link: VCF 4.3 Specification. Please note that variant files must contain at most 2000 variants.
Step 5: Choose to target SNPs individually or all within the 23-mer
To target all SNPs within the 23-mer sequence click the checkbox at the bottom of the form. This will design sgRNA's with all SNPs that overlap with the sequence rather than 1 sgRNA per SNP regardless of overlap.